Superfund Research Program
Three-Dimensional Cell Model Enhances DNA Damage Testing
View Research Brief as PDF(372KB)
Release Date: 06/03/2020
![]() subscribe/listen via iTunes, download(6.3MB), Transcript(101KB)
subscribe/listen via iTunes, download(6.3MB), Transcript(101KB)
NIEHS-funded Superfund Research Program (SRP) scientists developed a new platform, known as the SpheroidChip analysis method, to rapidly test for DNA damage in three-dimensional (3D) cell models. Development was led by Bevin Engelward, Sc.D., at the Massachusetts Institute of Technology.
DNA damage can increase risk of cancer and disease development. Scientists often study DNA damage and repair using cell cultures, and they are now using more physiologically relevant 3D cell culture models or spheroids. However, standard techniques for spheroid formation are technically challenging with relatively low throughput.
The new tool uses liver-cell spheroids to measure DNA damage in toxicity studies. Because the 3D aggregate structure allows cells to interact with each other, it more closely mimics a natural environment than testing with single cells. The new platform improves spheroid production; it is low cost, does not require specialized equipment, and can be assembled using basic tissue culture supplies and a reusable stamp for creating microwell patterns in agarose.
Creating a Spheroid Testing Platform
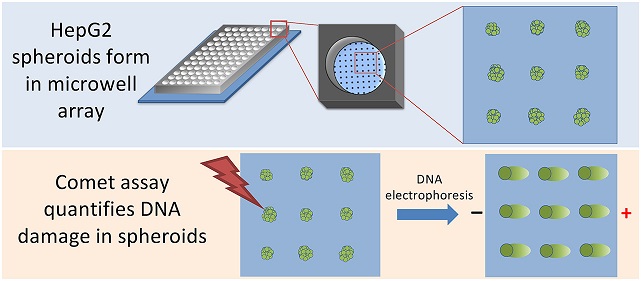 HepG2 spheroids form spontaneously in microwells. Once spheroids have formed, they can be immobilized on the chip with an agarose overlay. The researchers can then use the Comet assay to quantify DNA damage. (Image from Chao et al. 2020)
HepG2 spheroids form spontaneously in microwells. Once spheroids have formed, they can be immobilized on the chip with an agarose overlay. The researchers can then use the Comet assay to quantify DNA damage. (Image from Chao et al. 2020)Existing spheroid culture platforms create a non-adhesive environment that drives liver cells to aggregate into a spheroid. With these methods, only one spheroid can be cultured in each well. The research team developed a new technique using agarose, a nontoxic material suitable for cell growth that provides a low attachment environment and drives cell aggregation.
The researchers loaded human liver cancer cells, known as HepG2 cells, into 50 micrometer wide wells made in agarose. These microwells fit into the bottom of a standard well of a 96-well plate. They found that by using agarose to culture the cells, the HepG2 cells formed dozens of spheroids within each well in one to two days.
Assessing DNA Damage
They demonstrated that their system can analyze DNA damage levels, a novel application for spheroids. The approach leverages the CometChip assay to assess DNA damage directly in the agarose platform. The research team previously developed the CometChip assay, an adaptation of the standard laboratory Comet assay method that detects DNA strand breaks, but with higher throughput and better reproducibility.
The assay involves electrophoresis, or the movement of DNA under electric current through the agarose. When DNA is intact, it remains supercoiled and does not migrate significantly through the agarose during electrophoresis. In the presence of DNA damage, DNA is looser, which enables fragmented DNA to move through the agarose, producing a comet shape. Most of the DNA stays in place, which is the head of the comet, and the DNA that pulls away appears as a tail. The amount of DNA in the tail is proportional to the level of DNA strand breaks, which can be quantified through laboratory techniques and image analysis.
Intact HepG2 spheroids cultured in microwells can be electrophoresed to reveal the extent of DNA damage following exposure to inflammatory chemicals such as hydrogen peroxide. With this SpheroidChip analysis method, the researchers detected a dose-dependent increase in DNA damage and observed rapid repair of hydrogen peroxide-induced DNA damage.
DNA damage studies on intact spheroids were performed using steps similar to those in the CometChip experiments. Researchers treated spheroids with 0, 50, or 100 micromolar concentrations of hydrogen peroxide and measured increased migration of damaged DNA through agarose during electrophoresis, revealing the extent of DNA damage following exposure to hydrogen peroxide. (Image from Chao et al. 2020)
An Improved Method
One drawback of the traditional comet assay is that it requires single cells. As a result, cultured cells are broken up prior to being embedded in agarose and tested. Because DNA repair can be very rapid, significant DNA repair takes place during sample handling. In the case of hydrogen peroxide-induced DNA damage, repair is mostly complete by the time the samples are ready for comet analysis.
Using the SpheroidChip method, researchers can perform chemical exposures and the comet assay directly on the agarose chip with intact spheroids. This provides a distinct advantage for studying chemical exposures that lead to DNA damage that can then be rapidly repaired, including DNA damage created by inflammatory chemicals as well as certain chemical contaminants present at Superfund sites.
Using the agarose microarray, they were also able to condense what had previously required an entire 96-well plate into a single well, enabling new analysis techniques that were too cumbersome or impossible under conditions of a single spheroid per well.
For More Information Contact:
Bevin P. Engelward
Massachusetts Institute of Technology
77 Massachusetts Avenue
16-743C
Cambridge, Massachusetts 02139
Phone: 617-258-0260
Email: bevin@mit.edu
To learn more about this research, please refer to the following sources:
- Chao C, Ngo LP, Engelward BP. 2020. SpheroidChip: patterned agarose microwell compartments harboring HepG2 spheroids are compatible with genotoxicity testing. ACS Biomater Sci Eng 6:2427-2439. doi:10.1021/acsbiomaterials.9b01951
To receive monthly mailings of the Research Briefs, send your email address to srpinfo@niehs.nih.gov.


