Superfund Research Program
Using Fungi to Clean up Contaminated Soil
View Research Brief as PDF(694KB)
Release Date: 08/05/2020
![]() subscribe/listen via iTunes, download(5.7MB), Transcript(86KB)
subscribe/listen via iTunes, download(5.7MB), Transcript(86KB)
Native fungal communities point to a new way of cleaning up contaminated soil. After conducting a study to characterize fungi found in soil contaminated with polycyclic aromatic hydrocarbons (PAHs), researchers at the NIEHS-funded Superfund Research Program at Duke University discovered a group of fungi that may be promising for remediation.
 Researchers collected soil samples from the Holcomb Creosote Superfund Site in Yadkinville, North Carolina. (Image courtesy of Claudia Gunsch)
Researchers collected soil samples from the Holcomb Creosote Superfund Site in Yadkinville, North Carolina. (Image courtesy of Claudia Gunsch)Fungi are known to degrade a range of chemicals in the lab; however, they are not commonly used to break down chemicals at contaminated sites. Fungi added to contaminated soil often do not grow well in the presence of the contaminants and native bacteria.
Understanding the makeup of fungal communities that already thrive in contaminated soil may help tailor strategies for degrading contaminants, according to the study.
Duke researchers led by Claudia Gunsch, Ph.D., validated a strategy to detect existing onsite fungal communities that can grow in soil with high PAH levels. Native fungi that thrive with local bacteria and are more tolerant of PAHs may be able to metabolize the chemicals by producing degrading enzymes, which can detoxify PAH-polluted soils when their growth is stimulated.
Comparing PAH Levels and Fungal Communities
To study fungal communities in contaminated settings, the researchers collected soil samples from the Holcomb Creosote Superfund Site, a former wood treating facility. The soil from the site was contaminated with creosote, which was used as a wood preservative and contains a variety of PAHs, a large and widespread class of chemicals known for their carcinogenicity.
The researchers measured concentrations of 31 PAHs in soil from various locations at the Superfund site. The PAH concentrations varied widely, ranging from 307 nanograms per gram of soil to 26,196 nanograms per gram of soil. They also extracted and analyzed DNA in each sample to identify fungi present in the soil.
Even in the presence of higher levels of PAHs, the overall fungal diversity was not significantly affected, suggesting that a range of fungi are able to persist in the presence of high pollutant levels. When focusing on specific groups of fungi, they found that PAH tolerance was more common than PAH sensitivity in the fungal community.
They also identified PAH tolerant fungal groups that are closely related to known PAH degraders, suggesting these fungi may be good targets for stimulating the soil community to degrade PAHs. Additional fungal groups were positively associated with higher PAH levels but have not been previously characterized for PAH degradation, suggesting that they may also be good targets for further study.
Targeting Specific Fungi
The dominant fungi at the Holcomb Creosote site were ascomycetes, a large phylum commonly known as sac fungi.
Most fungal degradation strategies implemented in the field to date have focused on wood-rotting basidiomycete fungi because they produce powerful enzymes that can degrade many pollutants. In this study, researchers provide evidence that ascomycete may be more tolerant to high PAH concentrations than basidiomycetes under field conditions, suggesting that future treatment strategies may be more successful if they use ascomycetes.
According to the authors, the data suggest that PAH pollution acts as a selective pressure, shifting the community towards fungi that are resistant to PAHs, such as ascomycetes. That selective pressure of PAHs could improve the number of strains capable of metabolizing PAHs or otherwise degrading them.
In future work, the researchers suggest using genome sequencing methods to confirm the presence of specific degradation genes and demonstrate that the identified fungi contain genes responsible for carrying out specific degradation pathways.
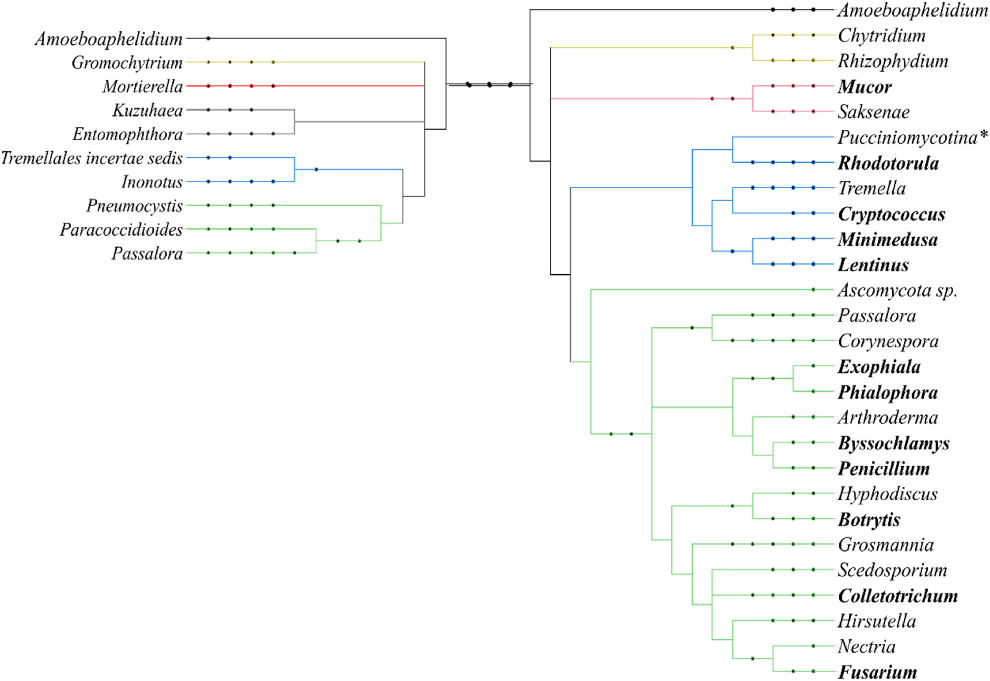 The researchers identified three times more genera that were positively correlated with PAH contamination, shown on the right, than genera that were negatively correlated, on the left. The highest number of positively correlated genera were ascomycete fungi, shown in green, followed basidiomycete fungi, displayed in blue. (Image from Czaplicki et al. 2020)
The researchers identified three times more genera that were positively correlated with PAH contamination, shown on the right, than genera that were negatively correlated, on the left. The highest number of positively correlated genera were ascomycete fungi, shown in green, followed basidiomycete fungi, displayed in blue. (Image from Czaplicki et al. 2020)For More Information Contact:
Claudia Gunsch
Duke University
Civil and Environmental Engineering
Box 90287
Durham, North Carolina 27708-0287
Phone: 919-660-5208
Email: ckgunsch@duke.edu
To learn more about this research, please refer to the following sources:
- Czaplicki LM, Redfern L, Cooper EM, Ferguson PL, Vilgalys R, Gunsch C. 2020. Investigating the mycobiome of the Holcomb Creosote Superfund Site. Chemosphere 252:9. doi:10.1016/j.chemosphere.2020.126208 PMID:32229362 PMCID:PMC7242165
To receive monthly mailings of the Research Briefs, send your email address to srpinfo@niehs.nih.gov.


